A ripple tells you something happened, but not exactly what.
That is the core problem behind a hard class of equations that scientists use when they try to work backward from what they can measure to what caused it. In weather systems, biology, and materials science, researchers often have the visible result, a shifting pattern, a temperature field, a cellular structure, but not the hidden rules or forces that produced it.
Engineers at the University of Pennsylvania say they have found a better way to tackle that problem with artificial intelligence. Their method, called “Mollifier Layers,” aims to make AI systems better at solving inverse partial differential equations, or inverse PDEs, especially when the data is noisy and the math gets unstable.
The work, published in Transactions on Machine Learning Research and set to be presented at NeurIPS 2026, offers a different route than the one much of modern AI has taken. Instead of leaning harder on larger models and more computing power, the Penn team turned to a mathematical idea that has been around for decades and reworked it for physics-informed machine learning.

“Solving an inverse problem is like looking at ripples in a pond and working backward to figure out where the pebble fell,” says Vivek Shenoy, Eduardo D. Glandt President’s Distinguished Professor in Materials Science and Engineering and senior author of the research. “You can see the effects clearly, but the real challenge is inferring the hidden cause.”
Differential equations are one of science’s basic tools for describing change. They can track how heat moves through a material, how a population grows, or how a chemical process unfolds. Partial differential equations go further, describing how those changes happen across both space and time.
Inverse PDEs flip the usual question. Instead of starting with known rules and predicting what will happen, they start with observations and ask what hidden parameters or dynamics must have been there in the first place.
That is a much harder task.
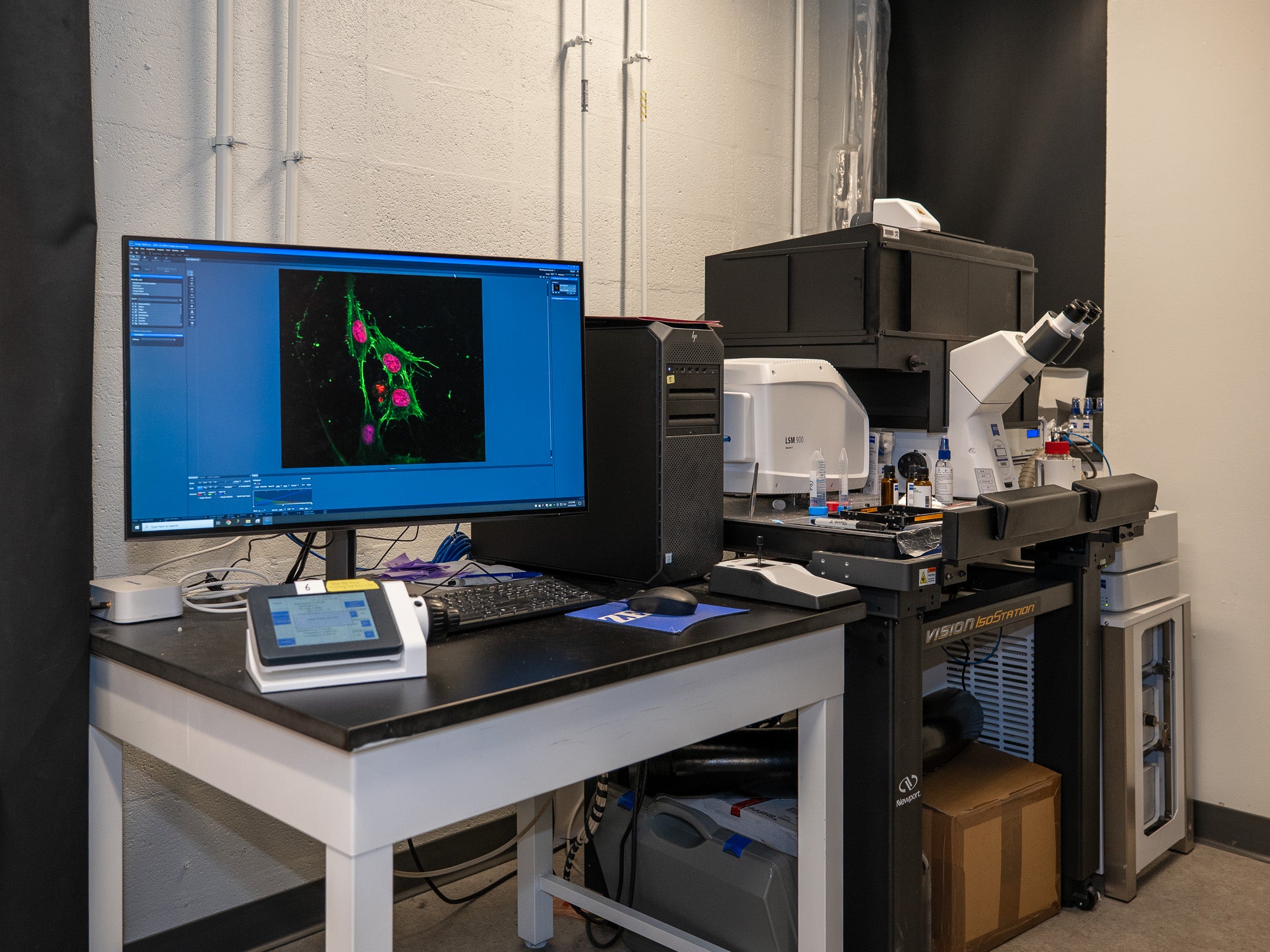
“For years, we’ve used these equations to study how chromatin, which is the folded state of DNA inside the nucleus, organizes itself inside living cells,” says Shenoy. “But we kept running into the same problem: We could see the structures and model their formation, but we could not reliably infer the epigenetic processes driving this system, namely the chemical changes that help control which genes are active. The more we tried to optimize the existing approach, the clearer it became that the mathematics itself needed to change.”
At the center of the bottleneck is differentiation, the math that measures how something changes. AI systems that handle inverse PDEs usually calculate derivatives through recursive automatic differentiation, a process that repeatedly traces how values change through a neural network.
That works, up to a point.

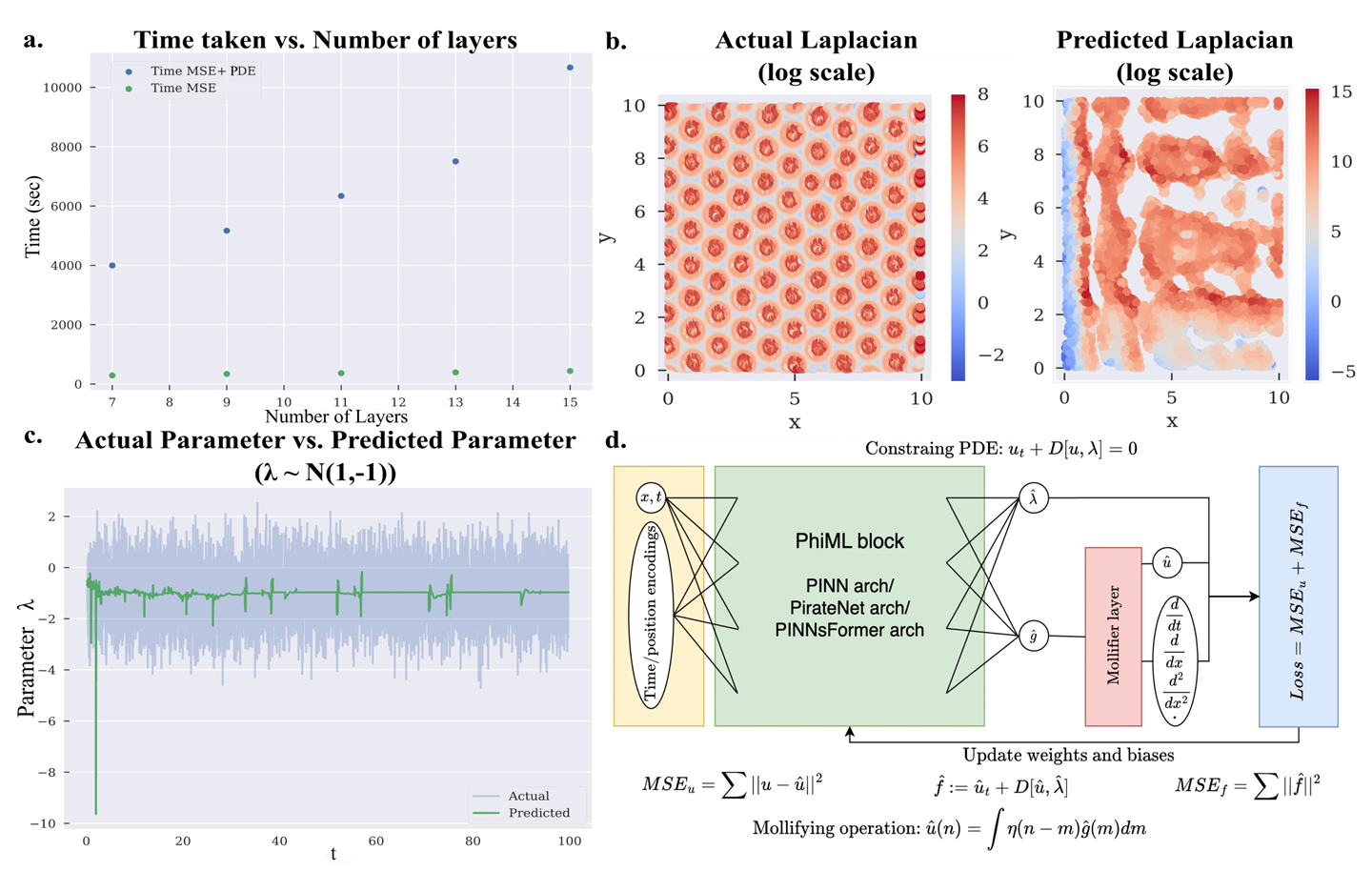
When the equations involve higher-order derivatives, and when the data contains noise, the process can become memory-hungry, slow, and unstable. The Penn researchers compared model behavior across several benchmark problems and found the strain rising sharply as systems became more complex. In one memory comparison using PINNs, or physics-informed neural networks, peak memory use climbed from 0.21 gigabytes to 2.70 gigabytes when a fourth-order reaction-diffusion problem used both data loss and PDE loss. Training time also ballooned as network depth increased.
The trouble is not just speed. Accuracy starts to wobble too. In a reaction-diffusion benchmark, the team reported that a PINN recovered the Laplacian with a correlation of only 0.21 against ground truth, a sign that the derivative estimate had drifted badly.
The researchers eventually decided the issue was not mainly the neural network design itself.
“We initially assumed the issue had to do with neural network’s architecture,” says Ananyae Kumar Bhartari, a graduate of Penn Engineering’s Scientific Computing master’s program and the other co-first author on the work. “But, after carefully adjusting the network, we eventually realized the bottleneck was recursive automatic differentiation itself.”
Their solution draws on mollifiers, a mathematical tool described in the 1940s by mathematician Kurt Otto Friedrichs. Mollifiers smooth jagged or noisy functions before you try to analyze them. Instead of taking unstable derivatives directly from the network’s output, the Penn system inserts a mollifier layer that smooths the signal first, then calculates derivatives through fixed convolution-based operations.
In plain terms, the method shifts the hardest part of the differentiation work away from repeated gradient calculations through the full network. The derivative comes from an analytic smoothing kernel instead.
That change appears to do several things at once. It reduces the memory burden, cuts training time, and makes the derivative estimates more stable under noise. The authors describe the layer as lightweight and architecture-agnostic, meaning it can be added to more than one kind of physics-informed machine learning model.
Vinayak Vinayak, a doctoral candidate in materials science and engineering and co-first author, puts the broader lesson simply: “Modern AI often advances by scaling up computation. But some scientific challenges require better mathematics, not just more compute.”
To see whether the approach held up, the team tested it on three classes of problems: a first-order 1D Langevin equation, a second-order 2D heat equation, and a fourth-order 2D reaction-diffusion system.
The last of those is especially punishing, because higher-order derivatives are where conventional methods often struggle most.
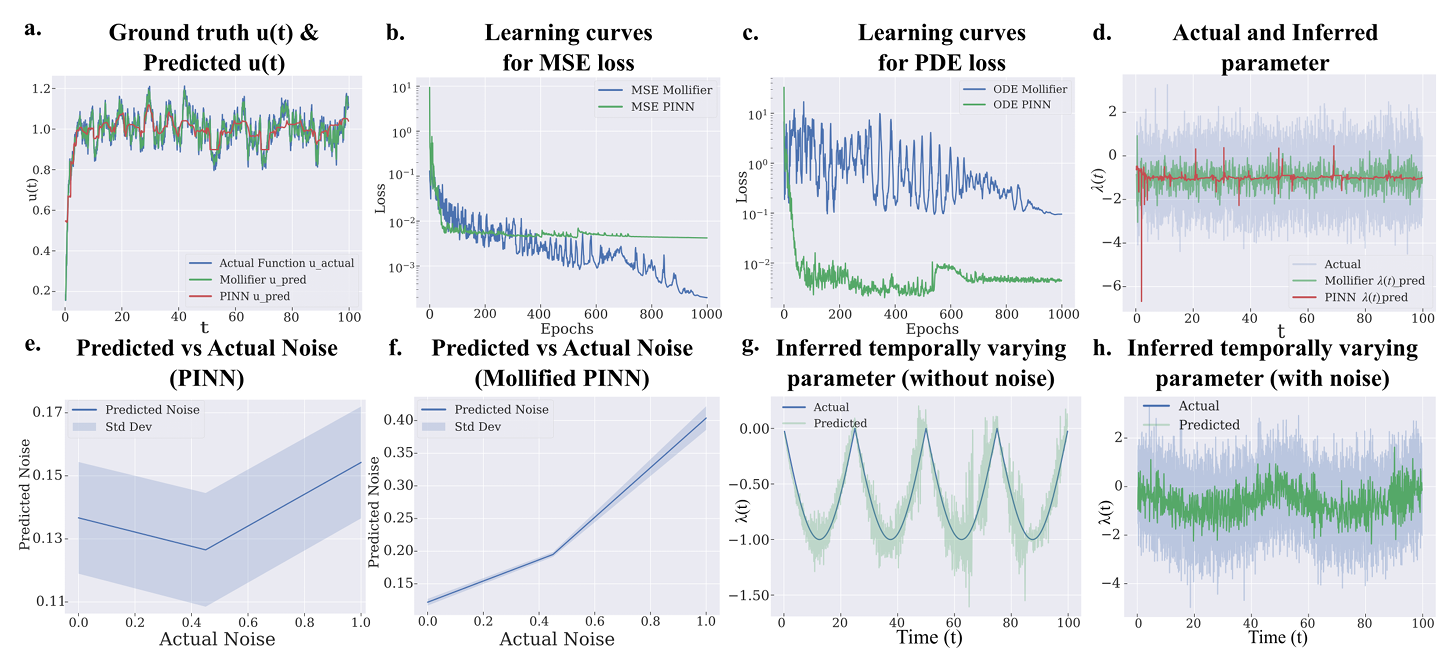
Across those benchmarks, mollifier-based models generally outperformed their native counterparts. In the first-order Langevin case, the mollified PINN achieved a temporal correlation of 0.97, compared with 0.36 for a standard PINN, while also using less memory, 0.16 gigabytes versus 0.21 gigabytes, and less training time, 1,615 seconds versus 2,138 seconds.
The gap widened in harder systems. For the second-order heat equation, the mollified PINN reached a spatial correlation of 0.99, while the standard PINN came in at 0.21. Peak memory use dropped from 1.20 gigabytes to 0.24 gigabytes. In the fourth-order reaction-diffusion benchmark, the same model cut training time from 3,386 seconds to 335 seconds and reduced peak memory from 2.75 gigabytes to 0.23 gigabytes. The mean correlation for the inferred parameter rose from 0.44 to 0.99.
The authors say the system reduced memory footprint and training time by about 6 to 10 times in their experiments.
Those gains mattered most when the hidden parameter changed across space or time, or when the signal carried noise. In those settings, mollified models were better at tracking the trends that standard systems often missed.
For Shenoy’s lab, the biological payoff comes from chromatin, the mix of DNA and proteins that packages chromosomes inside the nucleus.
Tiny chromatin domains, about 100 nanometers in size according to the source material, help regulate access to genetic material. That matters because accessibility influences gene expression, and gene expression shapes cell identity, function, aging, and disease.
“These domains are just 100 nanometers in size,” says Shenoy, “but because accessibility determines gene expression, and gene expression governs cell identity, function, aging and disease, these domains play a critical role in biology and health.”
The team had already been studying how epigenetic reactions and physical interactions organize chromatin structure. What they wanted was a more reliable way to infer the reaction rates behind those changes from what could actually be observed.
By doing that, researchers could move beyond snapshots of chromatin structure and toward models of how that structure changes over time. The work also extends to super-resolution microscopy, including STORM images of human cell nuclei, where the researchers say mollifier layers enabled spatially resolved inference of critical biophysical parameters from noisy image-derived fields.
In synthetic reaction-diffusion tests tied to DNA organization, the mollified PINNs recovered the spatially varying reaction rate with high accuracy and captured the Laplacian more faithfully than standard PINNs. The paper argues that this kind of parameter extraction could connect nanoscale chromatin remodeling to gene regulation, cancer metastasis, and cell fate memory in development and disease.
“If we can track how these reaction rates evolve during aging, cancer or development,” adds Vinayak, “this creates the potential for new therapies: If reaction rates control chromatin organization and cell fate, then altering those rates could redirect cells to desired states.”
The appeal of inverse PDEs is that they are not confined to one field.
Scientists run into them whenever they need to estimate hidden quantities such as diffusivity, conductance, or reaction rates from sparse or noisy measurements. That means the Penn framework could matter in areas far beyond chromatin. The source material points to materials science, fluid mechanics, genetics, and weather modeling as places where stable, efficient inference could help.
The authors also suggest the same principle might extend beyond inverse problems to forward models, operator learning, and neural ODE systems, all areas where accurate gradients matter.
Still, the work has limits.
Performance depends on the choice of mollifier kernel, which must balance noise suppression against the risk of washing out high-variance features. The current implementation also has weaknesses near boundaries and on anisotropic grids. The researchers say future work should explore adaptive or learned kernels, boundary-aware formulations, and validation strategies for adaptive meshes.
Those caveats matter because methods that look elegant on benchmarks do not always transfer cleanly to messier settings. Here, the researchers are clear that the mathematical shortcut helps, but does not erase the need for careful tuning.
This research points to a more efficient way for AI to extract hidden rules from difficult scientific data.
In practice, that could help researchers infer changing reaction rates in chromatin, recover spatially varying thermal properties in heat problems, or estimate unknown forcing terms in noisy dynamical systems. Because the method reduces memory use and training time while improving stability in higher-order problems, it may also make some physics-informed machine learning tasks more practical on real scientific datasets.
The broader value is not just speed. It is the possibility of turning observations into interpretable parameters with fewer failures when data is noisy or derivatives are hard to compute. As Shenoy puts it, “Ultimately, the goal is to move from observing complex patterns to quantitatively uncovering the rules that generate them. If you understand the rules that govern a system, you now have the possibility of changing it.”
Research findings are available online in the journal Transactions on Machine Learning Research.
The original story “Penn engineers use AI to solve some of science’s most difficult math problems” is published in The Brighter Side of News.
Like these kind of feel good stories? Get The Brighter Side of News’ newsletter.
The post Penn engineers use AI to solve some of science’s most difficult math problems appeared first on The Brighter Side of News.
Leave a comment
You must be logged in to post a comment.